Deep Learning for Genetic Variant Discovery
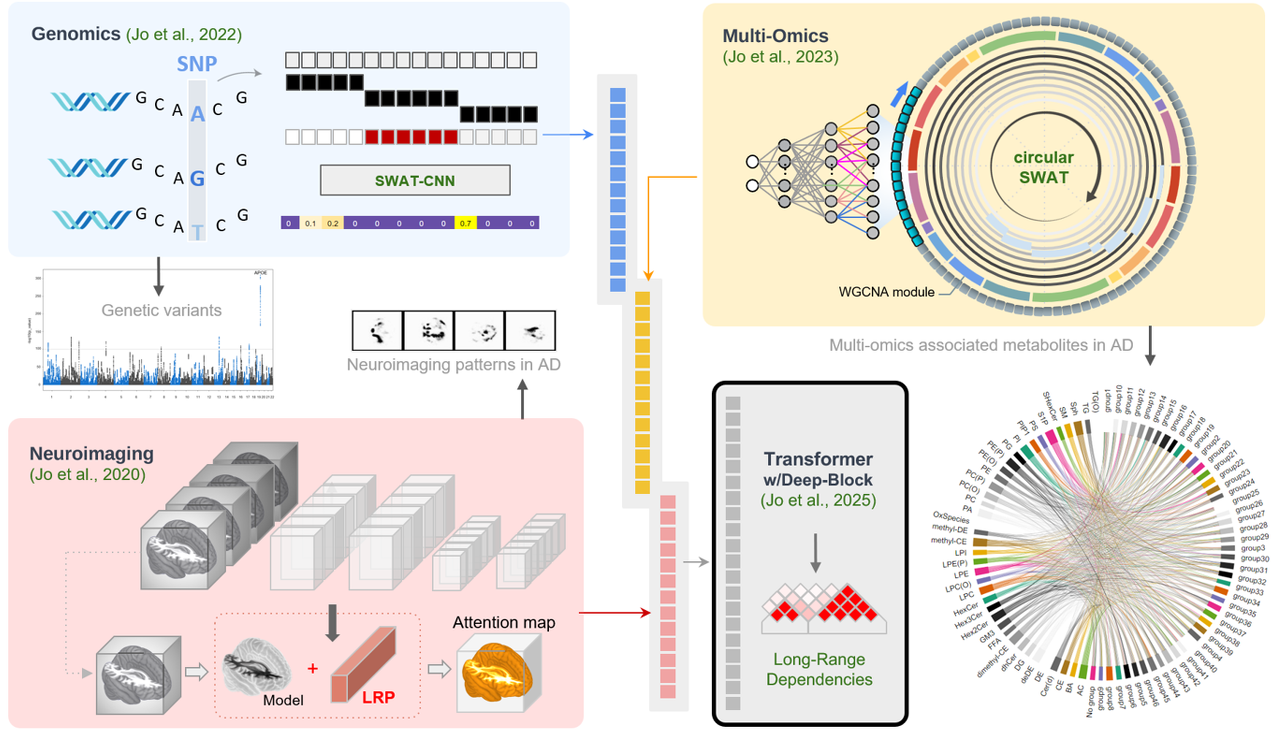
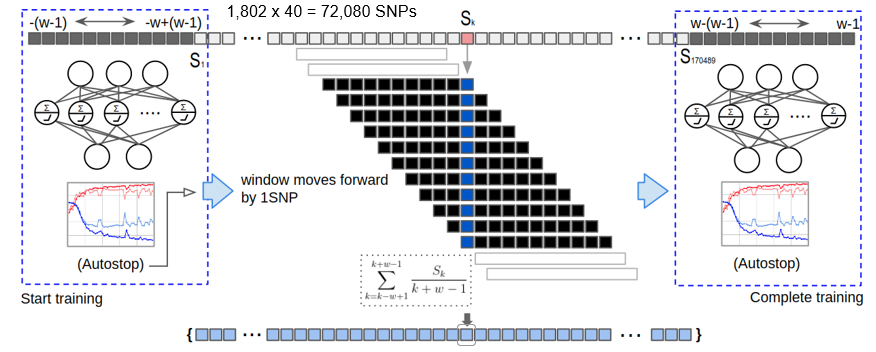
We develop deep learning frameworks to analyze whole-genome sequencing (WGS) data from large-scale cohorts such as the Alzheimer's Disease Sequencing Project (ADSP). Our SWAT-CNN approach applies a sliding-window strategy to detect AD-associated genetic variants, achieving an AUC of 0.82. More recently, our Deep-Block framework analyzed 7,416 WGS participants using linkage disequilibrium-informed deep learning, identifying novel SNPs beyond the well-known APOE region. We also developed DuAL-Net, a dual-attention architecture that integrates local genomic patterns with global biological knowledge for improved AD risk prediction from APOE-centered regional WGS data.
Selected Publications
- E.H. Lee, T. Jo*, DuAL-Net: A dual-network approach for Alzheimer's disease risk prediction using APOE-centered regional WGS data. Computational and Structural Biotechnology Journal, 2026.
- T. Jo* et al., LD-informed deep learning for Alzheimer's gene loci detection using WGS data. Alzheimer's & Dementia: TRCI, 2025.
- T. Jo et al., Deep learning-based identification of genetic variants: Application to Alzheimer's disease classification. Briefings in Bioinformatics, 2022.